Research in the Lab
Research in our lab is broadly focused on three parts, (1) Bioinformatics Analysis (2) DataBase Construction (3) Deep Learning for Medicine. Based on the platform of Wannan Medical College, we use bioinformatics big data and data analysis technology to conduct research on the molecular mechanism of diseases and develop related prediction tools.
Bioinformatics Analysis
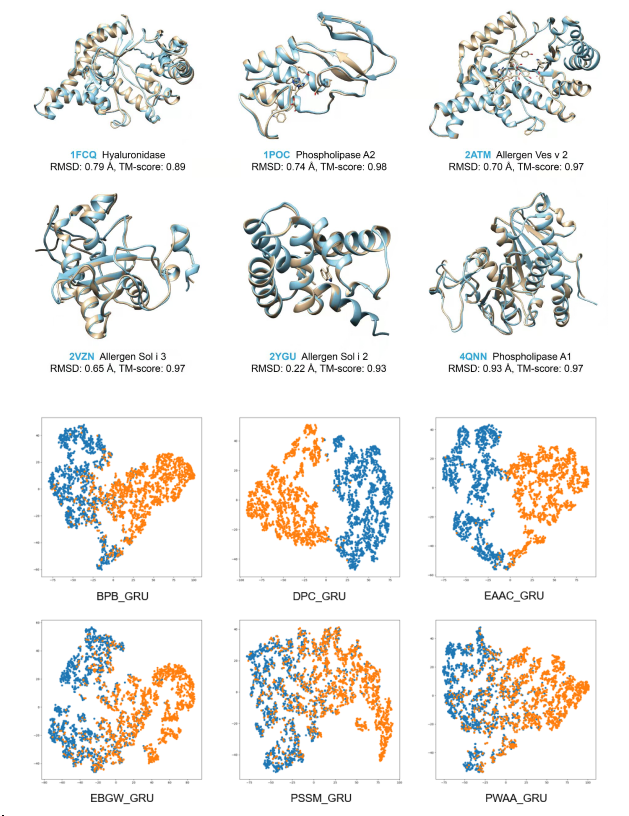
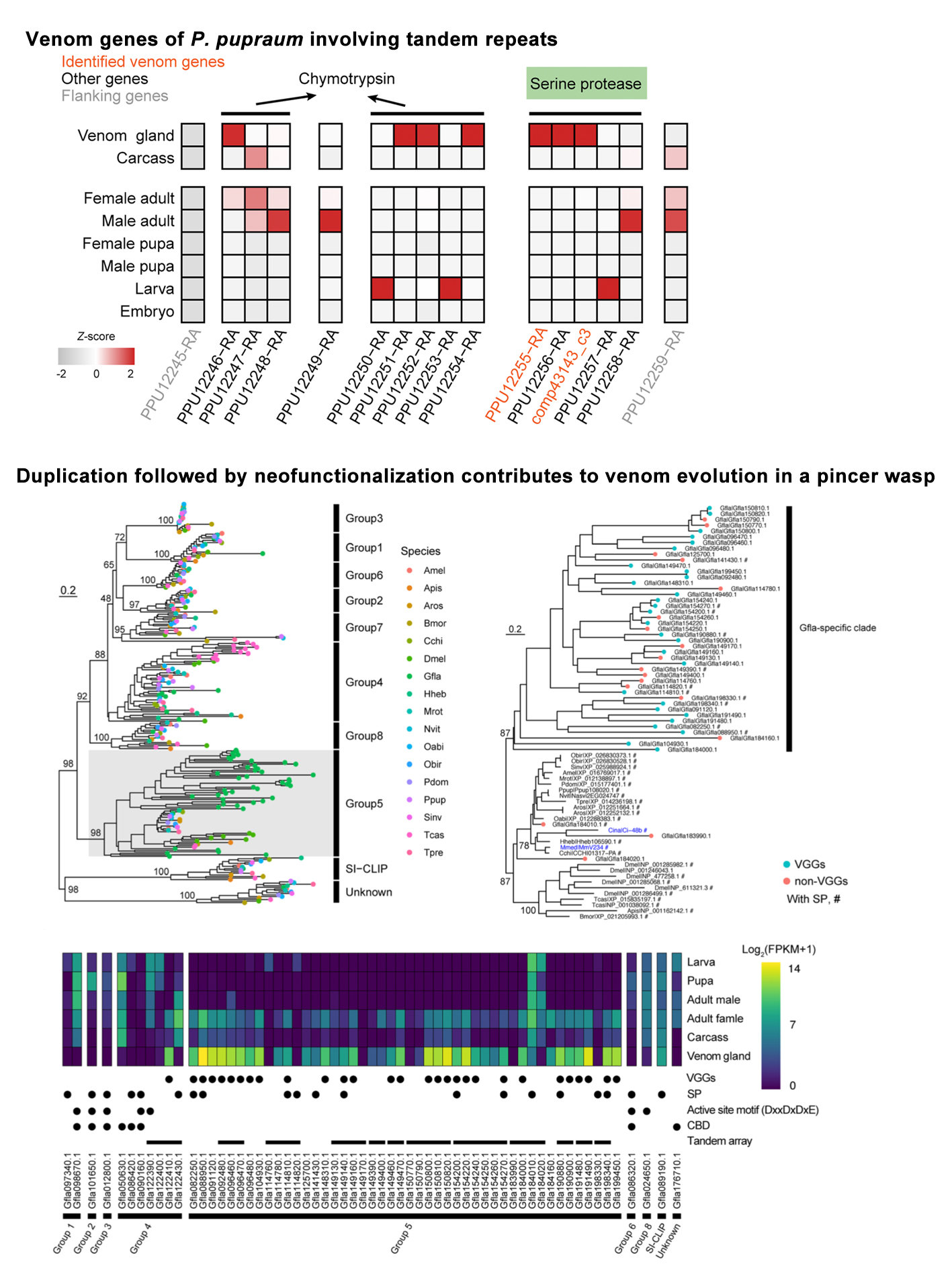
Venom system is a key model for studying evolutionary innovations. Parasitoid wasps use their venom as a major weapon against their hosts. To adapt to the varied hosts, parasitoid venom repertoires evolve rapidly, providing an excellent model for understanding how genes evolve to acquire their new “job” in venom. Using the high-quality data, we are currently studying the venom evolution of parasitoid wasps. Our results of many parasitoid wasps have showed that many different evolutionary models, including duplication followed by neofunctionalization, co-option, lateral gene transfer have involved in the venom evolution of parasitoid wasps. In the study of two Anastatus wasps, we find that the co-option evolution arose by expression shifts in the venom gland and plays a dominant role in venom turnover. In addition, this study uncovers the potential importance of transposable element insertions in driving gene expression shifts, contributing to their co-option in the venom gland. This discovery offers a reasonable explanation of the genetic basis underlying rapid venom turnover during parasitoid wasp evolution. Our ongoing project will strengthen this hypothesis through expanded sampling and functional studies. Overall, our works are important for understanding mechanisms of venom evolution and new gene function evolution. We have also developed a manually curated database, called iVenomDB (including over 4000 venom protein sequences of parasitoid wasps), to provide scientific community with the venom protein sequences, structures and other important functional information.
Database Construction
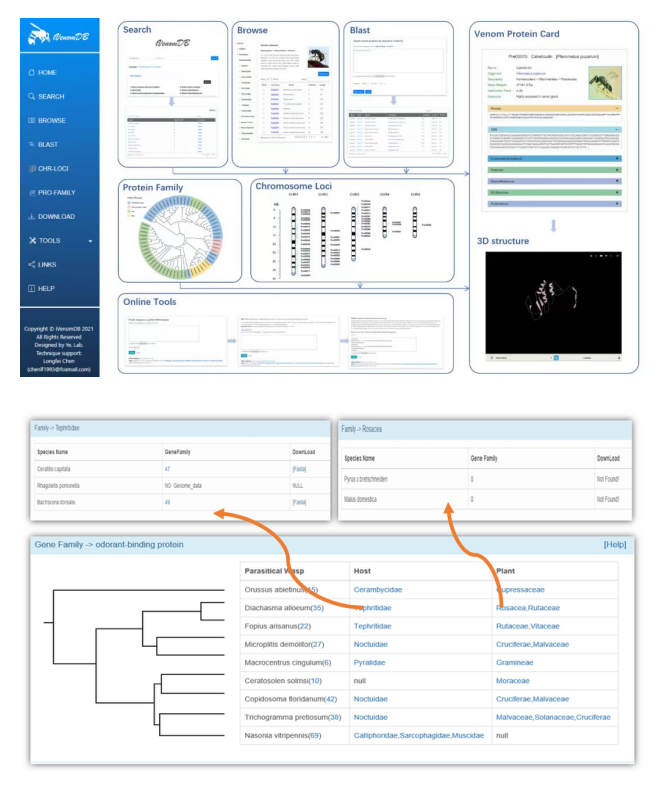
Bioinformatics database construction involves the organization, storage, and management of biological data in a structured format that enables efficient retrieval, analysis, and interpretation. These databases act as repositories of biological information, supporting research, analysis, and decision-making in the field of bioinformatics. It can provides researchers with easy access to a wide range of biological data for analysis and hypothesis testing; enables data sharing and collaboration across scientific communities, fostering new discoveries and insights; helps in studying genetic variations, drug interactions, disease mechanisms, and evolutionary relationships for informed decision-making in various fields. Our team developed several web databases, including iVenomDB, WaspBase and so on. iVenomDB fills a crucial gap in the field of bioinformatics by offering a specialized database dedicated to insect venom proteins.The manual curation process ensures high-quality data that can be used for in-depth analysis and research in the field of entomology and biotechnology. WaspBase offers a valuable resource for researchers aiming to unravel the complexities of interactions among parasitic wasps, insect hosts, and plants at the genomic level.This database facilitates data-driven studies on coevolutionary processes, host-parasite dynamics, and plant defense mechanisms, aiding in the development of sustainable pest management strategies and ecological conservation efforts.
Deep Learning for Biology
Bioinformatics algorithm and tools development refer to the process of creating computational methods and software applications specifically designed for analyzing biological data. These algorithms and tools are essential in tackling complex biological questions, interpreting genomic information, studying protein structures, and understanding biological processes at a molecular level. In bioinformatics, deep learning has emerged as a powerful tool for analyzing biological data, discovering patterns, and making predictions in areas such as genomics, proteomics, drug discovery, and personalized medicine. The content including (1) Genomic Sequencing Analysis: Deep learning models are used to analyze and interpret genomic sequences, identify genetic variations, predict gene function, and classify sequences based on structure and function. (2) Protein Structure Prediction: Deep learning algorithms can predict protein structures, infer protein-protein interactions, and classify protein functions based on sequence and structural information. (3) Drug Discovery: Deep learning is applied in virtual screening of drug compounds, predicting drug-target interactions, and optimizing drug design for more effective and targeted therapies. (4) Disease Diagnosis and Prediction: Deep learning models are used to analyze clinical data, medical images, and genetic information to diagnose diseases, predict disease progression, and stratify patients for personalized treatment.